Bioinformatic Profiling of Written Artefacts
2020–2025
RFF02
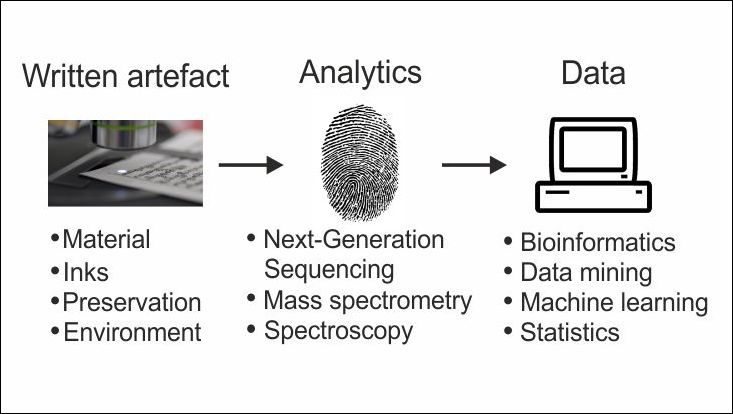
Our main activities in the cluster were in the context of the Palm-Leaf Manuscript Profiling Initiative (PLMPI), in which novel approaches for palm leaf manuscript profiling were developed and applied. The original sole focus was on the analysis of sequence data. However, the sampling of DNA and proteins was unfortunately not successful, which is why we adapted the scope and included other approaches. In close cooperation with the mobile lab of the CSMC we conducted chemometric analysis of data obtained by other analytical techniques, e.g. X-ray fluorescence (XRF), Fourier-Transform-Infrared (FTIR) and Raman spectroscopy. Our main results are therefore summarized in two areas: Chemometrics and Bioinformatics.
Bioinformatics: Our main focus here was on DNA analysis and we established specific pipelines for the analysis of the material and the metagenome (bacteria and other components that determine the environmental influences). In this context, we developed the approach GRADA (Grep Adapter analyser) for quality control of DNA data, which is especially important when ancient DNA is analysed. In addition, we generated a reference plastid sequence for the talipot palm. However, no approach could be developed to non-invasively sample DNA from the palm material. As we had no usable manuscript data available, we had to apply the approaches that we developed for genotyping [2, 4] and machine learning based feature selection and relation analysis [3] to other data sets to demonstrate their capabilities.
Chemometrics: We established fingerprint analysis of written artefacts with a handheld FTIR (DRIFTS) spectrometer (Exoscan) in combination with chemometrics and could show that the obtained data is reproducible, the treatment of the artefacts with oil has no significant influence on the spectra, and different artefacts can be distinguished [6]. In addition, we have been able to distinguish artefacts made from palms of different species, as well as artefacts of the same species from different geographical origins [6]. We have published the results of these analyses not only as a scientific paper, but also in RDR, along with an interactive visualisation of the results, making it easy for scientists from other disciplines to access and use the results [5]. We have also applied FTIR spectroscopy to the analysis of the 'Tsakalis' cards and have contributed a chapter to the book published in 2025 [7].
People
Project lead: Stephan Seifert
Research Associate: Lucas F. Voges
